
Phylogeny interpretation, types of trees, applications
A phylogeny, in evolutionary biology, it is a representation of the evolutionary history of a group of organisms or a species, emphasizing the line of descent and the kinship relationships between the groups.
Today, biologists have used data primarily from comparative morphology and anatomy, and from gene sequences to reconstruct thousands upon thousands of trees.
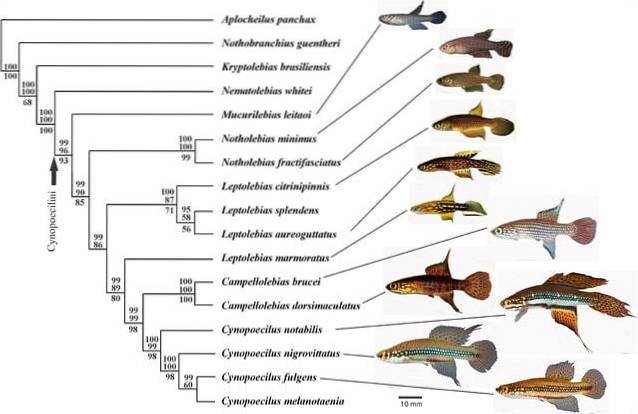
These trees seek to describe the evolutionary history of the different species of animals, plants, microbes and other organic beings that inhabit the earth..
The analogy with the tree of life dates from the time of Charles Darwin. This brilliant British naturalist captures in the masterpiece "The origin of species"A single image: a" tree "that represents the ramification of the lineages, starting from a common ancestor.
Article index
- 1 What is a phylogeny?
- 2 What is a phylogenetic tree?
- 3 How are phylogenetic trees interpreted?
- 4 How are phylogenies reconstructed?
- 4.1 Homologous characters
- 5 Types of trees
- 6 Politomias
- 7 Evolutionary classification
- 7.1 Monophyletic lineages
- 7.2 Paraphyletic and polyphyletic lineages
- 8 Applications
- 9 References
What is a phylogeny?
In the light of the biological sciences, one of the most amazing events that has taken place is evolution. This change in organic forms over time can be represented in a phylogenetic tree. Therefore, phylogeny expresses the history of lineages and how they have changed over time..
One of the direct implications of this graph is common ancestry. That is, all the organisms that we see today have emerged as descendants with modifications of past forms. This idea has been one of the most significant in the history of science.
All forms of life that we can appreciate today - from microscopic bacteria, to plants and the largest vertebrates - are connected and this relationship is represented in the vast and intricate tree of life..
Within the analogy of the tree, the species that live today would represent the leaves and the rest of the branches would be their evolutionary history.
What is a phylogenetic tree?
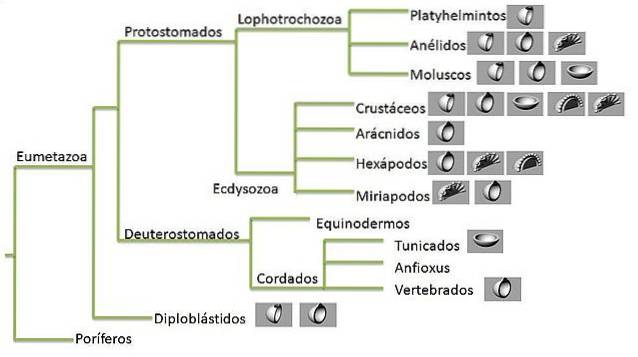
A phylogenetic tree is a graphic representation of the evolutionary history of a group of organisms. This pattern of historical relationships is the phylogeny that researchers are trying to estimate..
Trees consist of nodes that connect the "branches". The terminal nodes of each branch are the terminal taxa and represent the sequences or organisms for which data is known - these can be living or extinct species.
The internal nodes represent hypothetical ancestors, while the ancestor found at the root of the tree represents the ancestor of all the sequences represented in the graph..
How are phylogenetic trees interpreted?
There are many ways to represent a phylogenetic tree. For this reason, it is important to know how to recognize if these differences that are observed between two trees are due to different topologies - that is, real differences corresponding to two spellings - or are simply differences related to the style of the representation..
For example, the order in which the labels appear at the top can vary, without modifying the meaning of the graphic representation, generally the name of the species, genus, family, among other categories..
This occurs because the trees resemble a mobile, where the branches can rotate without changing the relationship of the represented species..
In this sense, it does not matter how many times the order is changed or the objects that are "hanging" are rotated, since it does not change the way in which they are connected - and that is the important thing..
How are phylogenies reconstructed?
Phylogenies are hypotheses that are formulated based on indirect evidence. Elucidating a phylogeny resembles the job of an investigator to solve a crime by following the clues from the crime scene.
Biologists often postulate their phylogenies using knowledge from various branches, such as paleontology, comparative anatomy, comparative embryology, and molecular biology..
The fossil record, although incomplete, provides very valuable information on the divergence times of groups of species.
With the passage of time, molecular biology has surpassed all the aforementioned fields, and most of the phylogenies are inferred from molecular data.
The goal of reconstructing a phylogenetic tree has a number of major drawbacks. There are approximately 1.8 million named species and many more without being described.
And, although a significant number of scientists strive every day to reconstruct relationships between species, there is still no complete tree..
Homologous characters
When biologists wish to describe the similarities between two structures or processes, they can do so in terms of common ancestry (homologies), analogies (function), or homoplasia (morphological resemblance)..
To reconstruct a phylogeny, exclusively homologous characters are used. Homology is a key concept in evolution and in the recreation of relationships between species, since only it adequately reflects the common ancestry of organisms.
Suppose we want to infer the phylogeny of three groups: birds, bats, and humans. To fulfill our objective, we decided to use the upper extremities as a characteristic that helps us discern the pattern of relationships..
Since birds and bats have modified structures for flight, we could erroneously conclude that bats and birds are more related to each other than bats to humans. Why have we come to the wrong conclusion? Because we have used an analogous and non-homologous character.
To find the correct relationship I must look for a homologous character, such as the presence of hair, mammary glands and three small bones in the middle ear - just to name a few. However, homologies are not easy to diagnose.
Types of trees
Not all trees are the same, there are different graphic representations and each one manages to incorporate some peculiar characteristic of the evolution of the group.
The most basic trees are cladograms. These graphs show the relationships in terms of common ancestry (according to the most recent common ancestors).
Additive trees contain additional information and are represented in the length of the branches.
The numbers associated with each branch correspond to some attribute in the sequence - such as the amount of evolutionary change that organisms have undergone. In addition to "additive trees", they are also known as metric trees or phylograms..
Ultrametric trees, also called dendograms, are a particular case of additive trees, where the tips of the tree are equidistant from the root to the tree.
These last two variants have all the data that we can find in a cladogram, and extra information. Therefore, they are not exclusive, if not complementary.
Politomias
Many times, the nodes of the trees are not fully resolved. Visually, it is said that there is a polytomy, when more than three branches emerge from a new one (there is a single ancestor for more than two immediate descendants). When a tree does not have polytomies, it is said to be fully resolved.
There are two types of polytomies. The first are the "hard" polytomies. These are intrinsic to the study group, and indicate that the descendants evolved at the same time. Alternatively, "soft" polytomies indicate unresolved relationships caused by data per se.
Evolutionary classification
Monophyletic lineages
Evolutionary biologists seek to find a classification that fits the branching pattern of the phylogenetic history of groups. In this process, a series of terms widely used in evolutionary biology have been developed: monophyletic, paraphyletic and polyphyletic..
A monophyletic taxon or lineage is one that comprises an ancestral species, which is represented in the node, and all its descendants, but not other species. This grouping is called a clade.
Monophyletic lineages are defined at each level of the taxonomic hierarchy. For example, the Family Felidae, a lineage that contains felines (including domestic cats), is considered monophyletic..
Similarly, Animalia is also a monophyletic taxon. As we see the family Felidae is within Animalia, so monophyletic groups can be nested.
Paraphyletic and polyphyletic lineages
However, not all biologists share cladistic classification thinking. In cases where the data are not complete or simply for convenience, certain taxa are named that include species from different clades or higher taxa that do not share a more recent common ancestor..
In this way, a taxon is polyphyletic, it is defined as a group that includes organisms from different clades, and these do not share a common ancestor. For example, if we want to designate a group of homeotherms, it would include birds and mammals..
In contrast, a paraphyletic group does not contain all the descendants of the most recent common ancestor. In other words, it excludes some of the members of the group. The most used example is reptiles, this group does not contain all the descendants of the most recent common ancestor: birds.
Applications
In addition to contributing to the tough task of elucidating the tree of life, phylogenies also have some fairly significant applications..
In the medical field, phylogenies are used to trace the origin and transmission rates of infectious diseases, such as AIDS, dengue fever, and influenza..
They are also used in the field of conservation biology. Knowledge of the phylogeny of an endangered species is essential to track the crossing patterns and the level of hybridization and inbreeding between individuals..
References
- Baum, D. A., Smith, S. D., & Donovan, S. S. (2005). The tree-thinking challenge. Science, 310(5750), 979-980.
- Curtis, H., & Barnes, N. S. (1994). Invitation to biology. Macmillan.
- Hall, B. K. (Ed.). (2012). Homology: The hierarchial basis of comparative biology. Academic Press.
- Hickman, C. P., Roberts, L. S., Larson, A., Ober, W. C., & Garrison, C. (2001). Integrated principles of zoology. McGraw-Hill.
- Hinchliff, CE, Smith, SA, Allman, JF, Burleigh, JG, Chaudhary, R., Coghill, LM, Crandall, KA, Deng, J., Drew, BT, Gazis, R., Gude, K., Hibbett, DS, Katz, LA, Laughinghouse, HD, McTavish, EJ, Midford, PE, Owen, CL, Ree, RH, Rees, JA, Soltis, DE, Williams, T.,… Cranston, KA (2015). Synthesis of phylogeny and taxonomy into a comprehensive tree of life. Proceedings of the National Academy of Sciences of the United States of America, 112(41), 12764-9.
- Kardong, K. V. (2006). Vertebrates: comparative anatomy, function, evolution. McGraw-Hill.
- Page, R. D., & Holmes, E. C. (2009). Molecular evolution: a phylogenetic approach. John Wiley & Sons.



Yet No Comments